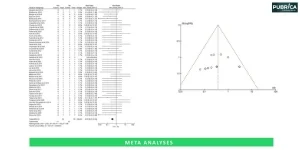
We use a mix of open-source tools (e.g., GATK, STAR, DESeq2, Cytoscape) and licensed platforms (e.g., Ingenuity Pathway Analysis, Partek). Each tool is carefully selected based on project objectives, dataset type, and peer-reviewed best practices, ensuring accuracy and reproducibility.
Our experts provide a free consultation to assess your research goals, data type (genomics, transcriptomics, proteomics, metabolomics, etc.), and publication requirements. Based on this, we recommend the most suitable bioinformatics package aligned with your needs and budget.
All workflows follow globally recognized best practices such as MIAME, MINSEQE, CONSORT, GCP/GLP, and FAIR data principles. This ensures that your results meet the highest scientific, ethical, and journal submission standards.
Yes. We specialize in integrative multi-omics analysis, combining molecular datasets with experimental or clinical data. This provides translational insights that support precision medicine, biomarker discovery, and real-world clinical applications.
Turnaround time depends on data size, complexity, and the chosen package. On average:
- Small datasets: 1–2 weeks
- Medium-scale projects: 3–4 weeks
- Large or multi-omics analyses: 5–8 weeks (priority delivery available)
We follow strict data security protocols. All client data is stored in encrypted systems, processed under HIPAA/GDPR-compliant frameworks, and never shared with third parties. NDAs (Non-Disclosure Agreements) can be signed on request.
Yes. Pubrica offers ongoing post-project support, including:
- Updates as new data becomes available
- Refinements to analyses based on reviewer feedback
- Additional visualizations or reports for publication and presentations
This ensures your results remain relevant, accurate, and publication-ready even after project delivery.